ANVI’O My First Bioinformatics Workshop Experience
This April, I had the chance to attend my first-ever bioinformatics workshop in Oldenburg, Germany, focused on Anvi’o an open‑source, community‑driven analysis and visualization platform for microbial ’omics and yes, that was as exciting (and terrifying) as it sounds. Oldenburg is a nice, tidy, and flat city. I personally found it extremely walkable, which was great because I got lost a few times but at least my legs survived. The workshop took place at the Helmholtz Institute for Functional Marine Biodiversity, and honestly, the building itself deserves its own paragraph.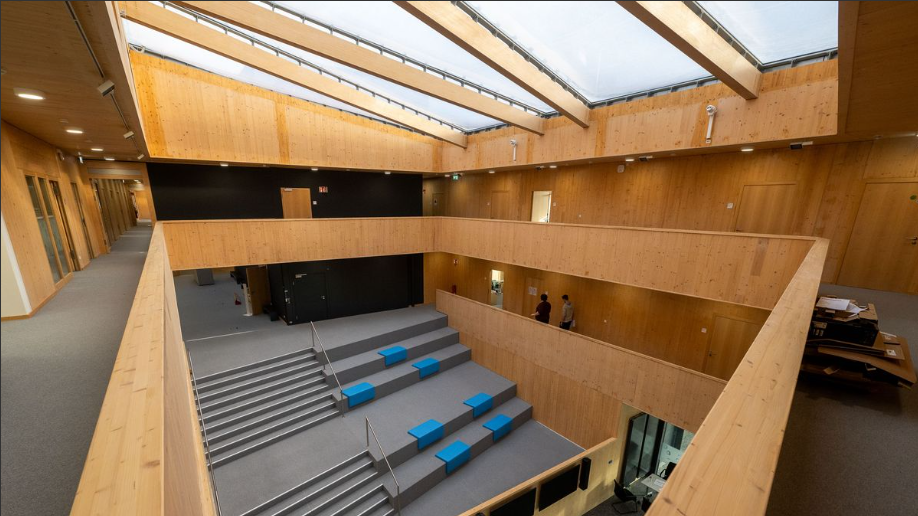  The building is made almost entirely of wood, and the ceiling is plastic because apparently wood can’t carry a heavy roof. My very first impression? A giant, very fancy sauna. Not complaining though it was warm, welcoming, and had great vibes, so I felt comfortable immediately.
The building is made almost entirely of wood, and the ceiling is plastic because apparently wood can’t carry a heavy roof. My very first impression? A giant, very fancy sauna. Not complaining though it was warm, welcoming, and had great vibes, so I felt comfortable immediately.
The workshop lasted five days. Knowledge-wise, it felt like five months were compressed into one week. But at the same time, when I was learning and coding, time passed so fast that I genuinely don’t understand how both of these feelings can exist together. Every day after lunch, we had a short symposium session, where participants gave 5-minute talks about their research or research interests. This was such a nice way to get to know everyone both scientifically and as actual humans.
The first day focused mainly on introducing the program:
- why it was developed,
- what problems it aims to solve,
- and how we can actually use it.
One of the coolest things is that the software is open source. You can help with the code, request new commands, or suggest improvements to make it easier to use.
For example, while learning interactive visualization, we noticed that if we wanted to “close” a parameter, we had to manually set its height to 0 so it wouldn’t appear. We all agreed this was not practical and asked for a simple “hide” button. The developers showed us where to submit feature requests, and one of the participants posted it.
And then, guess what?
The next day, the Anvi’o developers had already implemented it.
Honestly, still impressed.
Day 1 was mainly about genomics, setting the foundation for everything else. Lunch was amazing (perfect vegetarian food), and the drinks were organic German fizzy drinks. Since it was rhubarb season, they had rhubarb fizzy drinks, which instantly became my favourite.
The second day was all about phylogenetics. The trainers and Anvi’o developers started by explaining the concepts from scratch what phylogenetics is, why it matters, and how it connects to the analyses we were doing. Then we moved into tutorials using Anvi’o. This was my first time seriously working with terminal commands, and surprisingly… I understood them. They explained not only what commands do, but why they work that way, including what happens behind the scenes. That made it much easier for me to imagine applying the same steps to my own data.
While they explained each command, I caught myself thinking:
“If this were my dataset, how would I name it? How would I organise this?”
It felt like real preparation for my actual PhD work.
This day was also very special for another reason:
It was my first-ever PhD presentation. I talked about my research interests and my PhD topic something I had never presented before. I mentioned this in my talk because I was honestly nervous, but both the participants and trainers were incredibly kind and friendly. They made me feel like I had known them for years.
Maybe that’s also why five days felt like five months (in a good way).
The third day was more intense and my favourite day. First, they talked about metabolism how that section of Anvi’o was developed, what kinds of biological questions led to it, and what problems it aims to solve. Listening to the developers talk about their own tool development was incredibly inspiring. Their enthusiasm was contagious.
It made me feel like:
“Yes, I’m in the right place.”
Learning from people who genuinely love what they do and who can make others excited about it is something rare, especially outside academia.
From a technical side, we learned how Anvi’o predicts metabolism, which databases it uses, and ran tutorials where we generated metabolism tables using commands. The Wednesday lectures finished early, so I got the chance to walk around Oldenburg with some other participants. People were from Costa Rica, Canada, and many more places. I honestly love how science brings people together.
We explored the city while talking about research, cultures, and openly sharing which topics we found difficult. That was a huge relief for me I had secretly assumed everyone else understood everything. Realising that others were also confused sometimes gave me the confidence to ask more questions later.
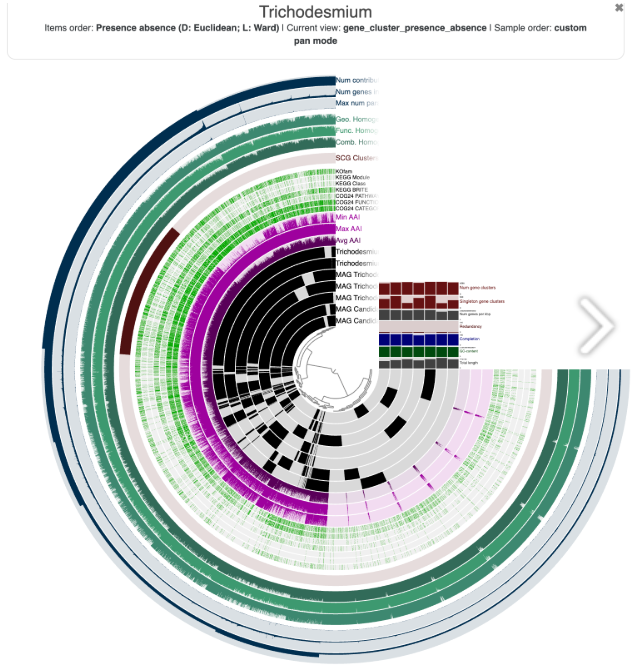 On the fourth day, we focused on pangenomics and more advanced visualisation techniques. This was when I really felt the power of visualisation complex datasets suddenly became something you could actually explore and interact with. It made me appreciate how important good visual design is, not just for analysis, but also for communication. The graph shown alongside highlights how much easier it is to understand the data when it’s presented visually.
On the fourth day, we focused on pangenomics and more advanced visualisation techniques. This was when I really felt the power of visualisation complex datasets suddenly became something you could actually explore and interact with. It made me appreciate how important good visual design is, not just for analysis, but also for communication. The graph shown alongside highlights how much easier it is to understand the data when it’s presented visually.
The fifth day moved into metagenomics, including discussions around read requirements and how analyses can be scaled up for larger datasets. This really highlighted how different working with real, complex datasets is compared to small test examples.
After listening to all the participant talks, I was struck by how diverse the research topics were, yet how they all connected through omics‑based microbial research. It was genuinely inspiring to see how different questions, systems, and scales connect through shared computational tools.
Overall, this workshop was an amazing experience scientifically, socially, and personally. I learned a lot, gained confidence, made new friends, and felt reassured that struggling while learning is not a weakness, but part of the process.

And yes… I still miss the rhubarb fizzy drinks and bakeries.
Esma Bozlak, PhD Student
Latest News
ANVI’O My First Bioinformatics Workshop Experience
This April, I had the chance to attend my first-ever bioinformatics workshop in Oldenburg, Germany, focused on Anvi’o an open‑source, community‑driven analysis and visualization platform for microbial